Capítulo 16 Disparidade e variação morfológica
Disparidade e variação são conceitos próximos, ambos relacionados à ideia de “variedade” ou “diversidade” morfológica; as mesmas métricas podem ser utilizadas para ambos. No entanto, uma distinção conceitual é aplicada: (i) Disparidade tem relação com a variedade de um grupo de espécies, e é o resultado do processo evolutivo de longo prazo; (ii) Variação tem relação com a variedade de indivíduos dentro de uma única população.
Vamos investigar a disparidade total e dentro/entre espécies de roedores sigmodontíneos.
require(geomorph)
#> Le chargement a nécessité le package : geomorph
#> Le chargement a nécessité le package : RRPP
#> Le chargement a nécessité le package : rgl
#> Le chargement a nécessité le package : Matrix
# Carregar arquivo tps com a vista ventral do crânio
tps.v<-readland.tps("dadosmg/ventral.dig_curso.pls.tps",specID = "ID", readcurves = FALSE)
# Gerar matriz com os pares de landmarks simétricos
pairs.matrix<-matrix(c(2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,24,22,28,23,29,19,25,20,26,21,27,30,31,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,55,56),nrow=26,ncol=2,byrow=T)
ind.v<-c(1:dim(tps.v)[3]) # vetor indivíduos
# GPA - Simetria bilateral - Extrair componente simetrico da forma
b.s<-bilat.symmetry(tps.v,ind=ind.v,object.sym=TRUE,land.pairs=pairs.matrix)
shape.v<-b.s$symm.shape # componente sim?trico da forma
ref.ventral<-mshape(shape.v)
# Carregar classificadores a partir de lista externa
plan<-read.table("dadosmg/Planilha_PLS.txt",h=T)
plan
species<-as.factor(plan[,2])
speciesDisparidade morfológica.
# Disparidade Morfológica
div.total<-morphol.disparity(shape.v~1,groups=NULL,iter=999)
#>
#> Please be aware that printing progress slows down the analysis (perhaps slightly).
#>
#> Preliminary Model Fit...
#>
#> Sums of Squares calculations: 3 permutations.
#>
|
| | 0%
|
|================= | 33%
|
|================================= | 67%
|
|==================================================| 100%
#>
#> No terms for ANOVA; only RSS calculated in each permutation
#> No factor in formula or model terms from which to define groups.
#> Procrustes variance:
#> 0.002195294
div.total
#> [1] 0.002195294
div.sp<-morphol.disparity(shape.v~species,groups=~species,iter=999)
#>
#> Please be aware that printing progress slows down the analysis (perhaps slightly).
#>
#> Preliminary Model Fit...
#>
#> Sums of Squares calculations: 3 permutations.
#>
|
| | 0%
|
|================= | 33%
|
|================================= | 67%
|
|==================================================| 100%
#>
#>
#> Performimg pairwise comparisons of disparity
#>
|
| | 0%
|
| | 1%
|
|= | 1%
|
|= | 2%
|
|= | 3%
|
|== | 3%
|
|== | 4%
|
|== | 5%
|
|=== | 5%
|
|=== | 6%
|
|=== | 7%
|
|==== | 7%
|
|==== | 8%
|
|==== | 9%
|
|===== | 9%
|
|===== | 10%
|
|===== | 11%
|
|====== | 11%
|
|====== | 12%
|
|====== | 13%
|
|======= | 13%
|
|======= | 14%
|
|======= | 15%
|
|======== | 15%
|
|======== | 16%
|
|======== | 17%
|
|========= | 17%
|
|========= | 18%
|
|========= | 19%
|
|========== | 19%
|
|========== | 20%
|
|========== | 21%
|
|=========== | 21%
|
|=========== | 22%
|
|=========== | 23%
|
|============ | 23%
|
|============ | 24%
|
|============ | 25%
|
|============= | 25%
|
|============= | 26%
|
|============= | 27%
|
|============== | 27%
|
|============== | 28%
|
|============== | 29%
|
|=============== | 29%
|
|=============== | 30%
|
|=============== | 31%
|
|================ | 31%
|
|================ | 32%
|
|================ | 33%
|
|================= | 33%
|
|================= | 34%
|
|================= | 35%
|
|================== | 35%
|
|================== | 36%
|
|================== | 37%
|
|=================== | 37%
|
|=================== | 38%
|
|=================== | 39%
|
|==================== | 39%
|
|==================== | 40%
|
|==================== | 41%
|
|===================== | 41%
|
|===================== | 42%
|
|===================== | 43%
|
|====================== | 43%
|
|====================== | 44%
|
|====================== | 45%
|
|======================= | 45%
|
|======================= | 46%
|
|======================= | 47%
|
|======================== | 47%
|
|======================== | 48%
|
|======================== | 49%
|
|========================= | 49%
|
|========================= | 50%
|
|========================= | 51%
|
|========================== | 51%
|
|========================== | 52%
|
|========================== | 53%
|
|=========================== | 53%
|
|=========================== | 54%
|
|=========================== | 55%
|
|============================ | 55%
|
|============================ | 56%
|
|============================ | 57%
|
|============================= | 57%
|
|============================= | 58%
|
|============================= | 59%
|
|============================== | 59%
|
|============================== | 60%
|
|============================== | 61%
|
|=============================== | 61%
|
|=============================== | 62%
|
|=============================== | 63%
|
|================================ | 63%
|
|================================ | 64%
|
|================================ | 65%
|
|================================= | 65%
|
|================================= | 66%
|
|================================= | 67%
|
|================================== | 67%
|
|================================== | 68%
|
|================================== | 69%
|
|=================================== | 69%
|
|=================================== | 70%
|
|=================================== | 71%
|
|==================================== | 71%
|
|==================================== | 72%
|
|==================================== | 73%
|
|===================================== | 73%
|
|===================================== | 74%
|
|===================================== | 75%
|
|====================================== | 75%
|
|====================================== | 76%
|
|====================================== | 77%
|
|======================================= | 77%
|
|======================================= | 78%
|
|======================================= | 79%
|
|======================================== | 79%
|
|======================================== | 80%
|
|======================================== | 81%
|
|========================================= | 81%
|
|========================================= | 82%
|
|========================================= | 83%
|
|========================================== | 83%
|
|========================================== | 84%
|
|========================================== | 85%
|
|=========================================== | 85%
|
|=========================================== | 86%
|
|=========================================== | 87%
|
|============================================ | 87%
|
|============================================ | 88%
|
|============================================ | 89%
|
|============================================= | 89%
|
|============================================= | 90%
|
|============================================= | 91%
|
|============================================== | 91%
|
|============================================== | 92%
|
|============================================== | 93%
|
|=============================================== | 93%
|
|=============================================== | 94%
|
|=============================================== | 95%
|
|================================================ | 95%
|
|================================================ | 96%
|
|================================================ | 97%
|
|================================================= | 97%
|
|================================================= | 98%
|
|================================================= | 99%
|
|==================================================| 99%
|
|==================================================| 100%
div.sp
#>
#> Call:
#> morphol.disparity(f1 = shape.v ~ species, groups = ~species,
#> iter = 999)
#>
#>
#>
#> Randomized Residual Permutation Procedure Used
#> 1000 Permutations
#>
#> Procrustes variances for defined groups
#> Aded Aega Aexa Aupi
#> 0.0004630470 0.0002893775 0.0006065943 0.0003949857
#> Elmo Irta Jshu Lomi
#> 0.0001739802 0.0003795813 0.0004866557 0.0003683766
#>
#>
#> Pairwise absolute differences between variances
#> Aded Aega Aexa Aupi
#> Aded 0.000000e+00 1.736695e-04 0.0001435473 6.806128e-05
#> Aega 1.736695e-04 0.000000e+00 0.0003172168 1.056082e-04
#> Aexa 1.435473e-04 3.172168e-04 0.0000000000 2.116086e-04
#> Aupi 6.806128e-05 1.056082e-04 0.0002116086 0.000000e+00
#> Elmo 2.890668e-04 1.153973e-04 0.0004326141 2.210055e-04
#> Irta 8.346564e-05 9.020385e-05 0.0002270129 1.540436e-05
#> Jshu 2.360875e-05 1.972782e-04 0.0001199386 9.167002e-05
#> Lomi 9.467042e-05 7.899906e-05 0.0002382177 2.660915e-05
#> Elmo Irta Jshu Lomi
#> Aded 0.0002890668 8.346564e-05 2.360875e-05 9.467042e-05
#> Aega 0.0001153973 9.020385e-05 1.972782e-04 7.899906e-05
#> Aexa 0.0004326141 2.270129e-04 1.199386e-04 2.382177e-04
#> Aupi 0.0002210055 1.540436e-05 9.167002e-05 2.660915e-05
#> Elmo 0.0000000000 2.056012e-04 3.126755e-04 1.943964e-04
#> Irta 0.0002056012 0.000000e+00 1.070744e-04 1.120479e-05
#> Jshu 0.0003126755 1.070744e-04 0.000000e+00 1.182792e-04
#> Lomi 0.0001943964 1.120479e-05 1.182792e-04 0.000000e+00
#>
#>
#> P-Values
#> Aded Aega Aexa Aupi Elmo Irta Jshu Lomi
#> Aded 1.000 0.175 0.247 0.600 0.021 0.520 0.844 0.468
#> Aega 0.175 1.000 0.009 0.407 0.343 0.477 0.118 0.527
#> Aexa 0.247 0.009 1.000 0.089 0.001 0.062 0.353 0.058
#> Aupi 0.600 0.407 0.089 1.000 0.072 0.917 0.469 0.842
#> Elmo 0.021 0.343 0.001 0.072 1.000 0.097 0.011 0.146
#> Irta 0.520 0.477 0.062 0.917 0.097 1.000 0.418 0.934
#> Jshu 0.844 0.118 0.353 0.469 0.011 0.418 1.000 0.361
#> Lomi 0.468 0.527 0.058 0.842 0.146 0.934 0.361 1.000
# Comparar com dispersão de pontos nos primeiros eixos da PCA:
col.group<-rainbow(length(levels(species))) # criar vetor de cores para grupos
names(col.group)<-levels(species)
col.group<-col.group[match(species,names(col.group))]
PCA<-gm.prcomp(shape.v) # rodar a PCA
plot(PCA$x[,1],PCA$x[,2],pch=21,cex=2,bg=col.group,asp=T)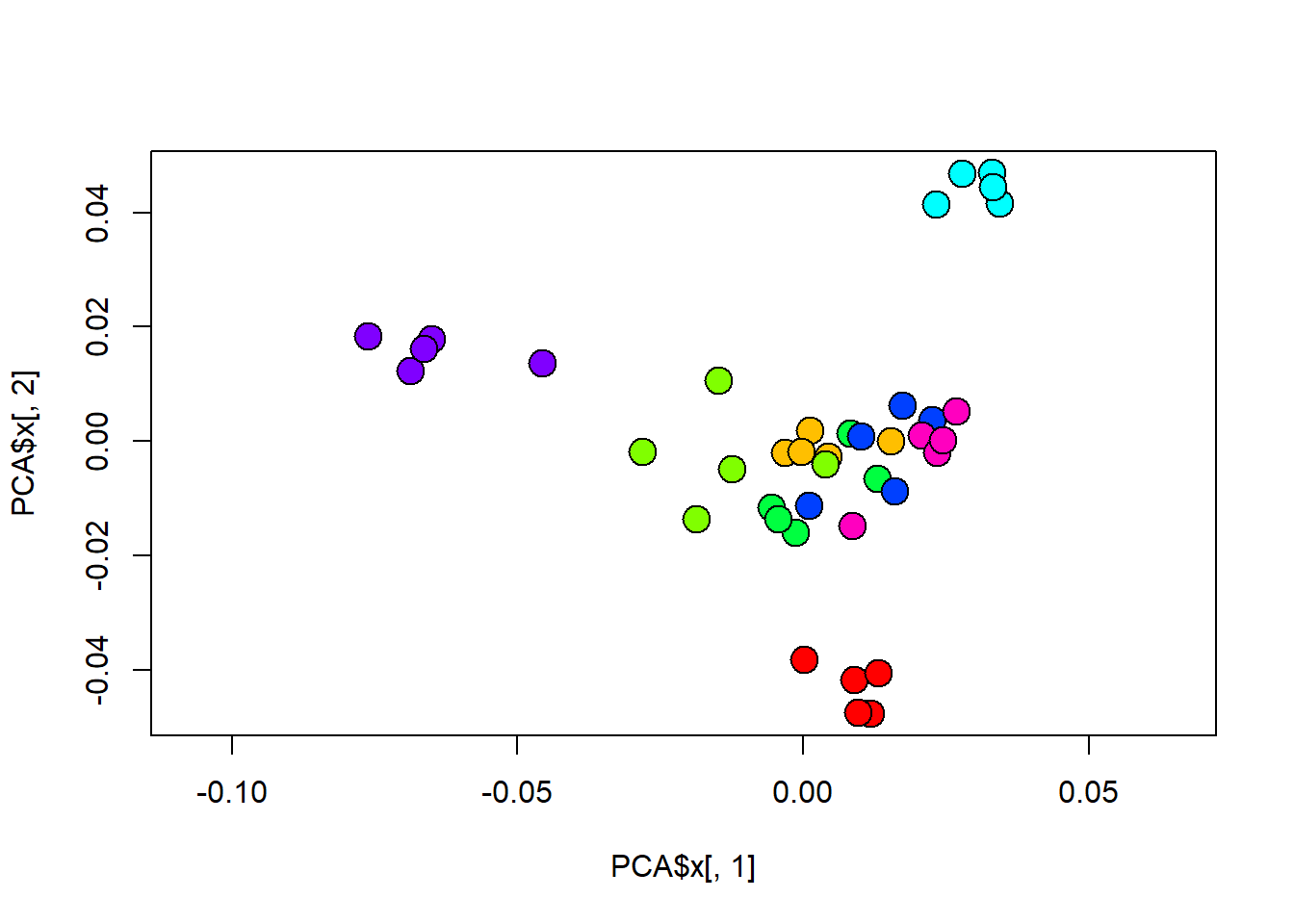
# Disparidade considerando alometria
gpa.v<-gpagen(tps.v)
#>
#> Performing GPA
#>
|
| | 0%
|
|============ | 25%
|
|========================= | 50%
|
|==================================================| 100%
#>
#> Making projections... Finished!
size<-gpa.v$Csize
div.sp<-morphol.disparity(shape.v~log(size)*species,groups=~species,iter=999)
#>
#> Please be aware that printing progress slows down the analysis (perhaps slightly).
#>
#> Preliminary Model Fit...
#>
#> Sums of Squares calculations: 3 permutations.
#>
|
| | 0%
|
|================= | 33%
|
|================================= | 67%
|
|==================================================| 100%
#>
#>
#> Performimg pairwise comparisons of disparity
#>
|
| | 0%
|
| | 1%
|
|= | 1%
|
|= | 2%
|
|= | 3%
|
|== | 3%
|
|== | 4%
|
|== | 5%
|
|=== | 5%
|
|=== | 6%
|
|=== | 7%
|
|==== | 7%
|
|==== | 8%
|
|==== | 9%
|
|===== | 9%
|
|===== | 10%
|
|===== | 11%
|
|====== | 11%
|
|====== | 12%
|
|====== | 13%
|
|======= | 13%
|
|======= | 14%
|
|======= | 15%
|
|======== | 15%
|
|======== | 16%
|
|======== | 17%
|
|========= | 17%
|
|========= | 18%
|
|========= | 19%
|
|========== | 19%
|
|========== | 20%
|
|========== | 21%
|
|=========== | 21%
|
|=========== | 22%
|
|=========== | 23%
|
|============ | 23%
|
|============ | 24%
|
|============ | 25%
|
|============= | 25%
|
|============= | 26%
|
|============= | 27%
|
|============== | 27%
|
|============== | 28%
|
|============== | 29%
|
|=============== | 29%
|
|=============== | 30%
|
|=============== | 31%
|
|================ | 31%
|
|================ | 32%
|
|================ | 33%
|
|================= | 33%
|
|================= | 34%
|
|================= | 35%
|
|================== | 35%
|
|================== | 36%
|
|================== | 37%
|
|=================== | 37%
|
|=================== | 38%
|
|=================== | 39%
|
|==================== | 39%
|
|==================== | 40%
|
|==================== | 41%
|
|===================== | 41%
|
|===================== | 42%
|
|===================== | 43%
|
|====================== | 43%
|
|====================== | 44%
|
|====================== | 45%
|
|======================= | 45%
|
|======================= | 46%
|
|======================= | 47%
|
|======================== | 47%
|
|======================== | 48%
|
|======================== | 49%
|
|========================= | 49%
|
|========================= | 50%
|
|========================= | 51%
|
|========================== | 51%
|
|========================== | 52%
|
|========================== | 53%
|
|=========================== | 53%
|
|=========================== | 54%
|
|=========================== | 55%
|
|============================ | 55%
|
|============================ | 56%
|
|============================ | 57%
|
|============================= | 57%
|
|============================= | 58%
|
|============================= | 59%
|
|============================== | 59%
|
|============================== | 60%
|
|============================== | 61%
|
|=============================== | 61%
|
|=============================== | 62%
|
|=============================== | 63%
|
|================================ | 63%
|
|================================ | 64%
|
|================================ | 65%
|
|================================= | 65%
|
|================================= | 66%
|
|================================= | 67%
|
|================================== | 67%
|
|================================== | 68%
|
|================================== | 69%
|
|=================================== | 69%
|
|=================================== | 70%
|
|=================================== | 71%
|
|==================================== | 71%
|
|==================================== | 72%
|
|==================================== | 73%
|
|===================================== | 73%
|
|===================================== | 74%
|
|===================================== | 75%
|
|====================================== | 75%
|
|====================================== | 76%
|
|====================================== | 77%
|
|======================================= | 77%
|
|======================================= | 78%
|
|======================================= | 79%
|
|======================================== | 79%
|
|======================================== | 80%
|
|======================================== | 81%
|
|========================================= | 81%
|
|========================================= | 82%
|
|========================================= | 83%
|
|========================================== | 83%
|
|========================================== | 84%
|
|========================================== | 85%
|
|=========================================== | 85%
|
|=========================================== | 86%
|
|=========================================== | 87%
|
|============================================ | 87%
|
|============================================ | 88%
|
|============================================ | 89%
|
|============================================= | 89%
|
|============================================= | 90%
|
|============================================= | 91%
|
|============================================== | 91%
|
|============================================== | 92%
|
|============================================== | 93%
|
|=============================================== | 93%
|
|=============================================== | 94%
|
|=============================================== | 95%
|
|================================================ | 95%
|
|================================================ | 96%
|
|================================================ | 97%
|
|================================================= | 97%
|
|================================================= | 98%
|
|================================================= | 99%
|
|==================================================| 99%
|
|==================================================| 100%
div.sp
#>
#> Call:
#> morphol.disparity(f1 = shape.v ~ log(size) * species, groups = ~species,
#> iter = 999)
#>
#>
#>
#> Randomized Residual Permutation Procedure Used
#> 1000 Permutations
#>
#> Procrustes variances for defined groups
#> Aded Aega Aexa Aupi
#> 0.0003260927 0.0001331069 0.0004333711 0.0001910877
#> Elmo Irta Jshu Lomi
#> 0.0001310704 0.0002895081 0.0003230759 0.0002042844
#>
#>
#> Pairwise absolute differences between variances
#> Aded Aega Aexa Aupi
#> Aded 0.000000e+00 1.929858e-04 0.0001072784 1.350051e-04
#> Aega 1.929858e-04 0.000000e+00 0.0003002643 5.798077e-05
#> Aexa 1.072784e-04 3.002643e-04 0.0000000000 2.422835e-04
#> Aupi 1.350051e-04 5.798077e-05 0.0002422835 0.000000e+00
#> Elmo 1.950223e-04 2.036498e-06 0.0003023008 6.001727e-05
#> Irta 3.658457e-05 1.564013e-04 0.0001438630 9.842049e-05
#> Jshu 3.016813e-06 1.899690e-04 0.0001102952 1.319882e-04
#> Lomi 1.218084e-04 7.117747e-05 0.0002290868 1.319670e-05
#> Elmo Irta Jshu Lomi
#> Aded 1.950223e-04 3.658457e-05 3.016813e-06 1.218084e-04
#> Aega 2.036498e-06 1.564013e-04 1.899690e-04 7.117747e-05
#> Aexa 3.023008e-04 1.438630e-04 1.102952e-04 2.290868e-04
#> Aupi 6.001727e-05 9.842049e-05 1.319882e-04 1.319670e-05
#> Elmo 0.000000e+00 1.584378e-04 1.920055e-04 7.321397e-05
#> Irta 1.584378e-04 0.000000e+00 3.356776e-05 8.522379e-05
#> Jshu 1.920055e-04 3.356776e-05 0.000000e+00 1.187915e-04
#> Lomi 7.321397e-05 8.522379e-05 1.187915e-04 0.000000e+00
#>
#>
#> P-Values
#> Aded Aega Aexa Aupi Elmo Irta Jshu Lomi
#> Aded 1.000 0.096 0.350 0.241 0.098 0.760 0.980 0.306
#> Aega 0.096 1.000 0.012 0.614 0.980 0.162 0.104 0.521
#> Aexa 0.350 0.012 1.000 0.043 0.009 0.252 0.331 0.043
#> Aupi 0.241 0.614 0.043 1.000 0.581 0.398 0.236 0.915
#> Elmo 0.098 0.980 0.009 0.581 1.000 0.158 0.092 0.527
#> Irta 0.760 0.162 0.252 0.398 0.158 1.000 0.760 0.474
#> Jshu 0.980 0.104 0.331 0.236 0.092 0.760 1.000 0.302
#> Lomi 0.306 0.521 0.043 0.915 0.527 0.474 0.302 1.000